|
Addgene inc
addgene plasmid 122091 Addgene Plasmid 122091, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/addgene plasmid 122091/product/Addgene inc Average 93 stars, based on 1 article reviews
addgene plasmid 122091 - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
Addgene inc
pbluescript sgrna expression plasmids 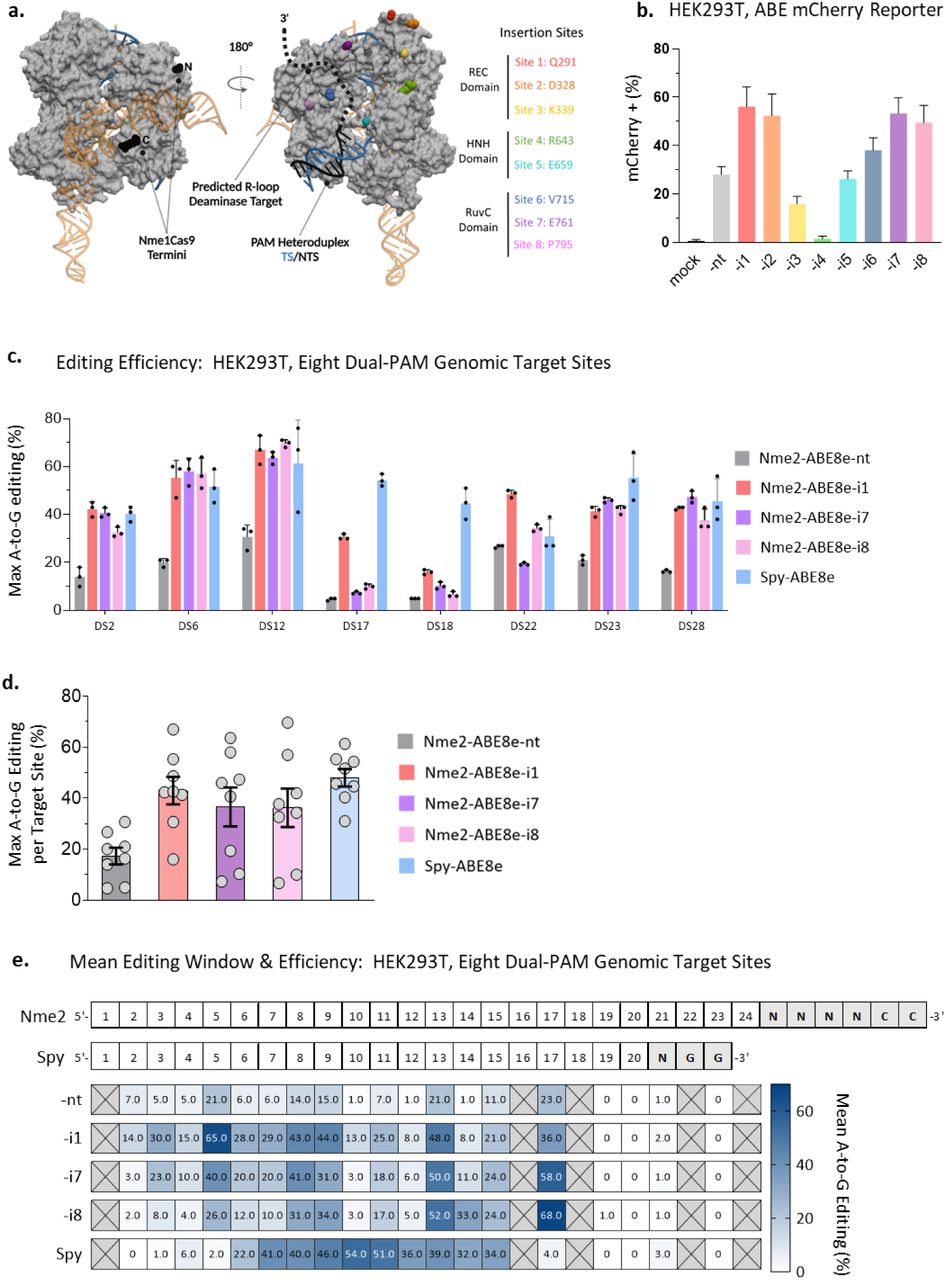 Pbluescript Sgrna Expression Plasmids, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pbluescript sgrna expression plasmids/product/Addgene inc Average 93 stars, based on 1 article reviews
pbluescript sgrna expression plasmids - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
Image Search Results

Journal: bioRxiv
Article Title: Engineering Nme2Cas9 Adenine Base Editors with Improved Activity and Targeting Scope
doi: 10.1101/2023.04.14.536905
Figure Lengend Snippet: a) Nme1Cas9/sgRNA/DNA ternary complex structure, PDB:6JDV. Nme2Cas9 is 98% identical to Nme2Cas9 outside of the WED and PAM-interacting domains. Black spheres represent N-and C-termini and colored spheres represent sites of domain insertion. Deaminase domain insertion sites (Nme2Cas9 aa numbers) are specified to the right, with colors matching the sites indicated in the structure. b) Activities of Nme2-ABE8e constructs in mCherry reporter cells (activated upon A-to-G editing) after plasmid transfection, measured by flow cytometry (n = 3 biological replicates in technical duplicate; data represent mean ± SD). c) A-to-G editing following transfection of Spy-ABE8e vs. Nme2-ABE8e plasmids, using PAM-matched, endogenous HEK293T genomic loci. The editing efficiency at the maximally edited adenine for each target was plotted. Editing efficiencies were measured by amplicon deep sequencing (n = 3 biological replicates; data represent mean ± SD). d) Data from (c) were aggregated and replotted, with each data point representing the maximum A-to-G editing efficiency of an individual target site, as measured by amplicon deep sequencing (n = 3 biological replicates; data represent mean ± SEM). e) Summary of mean A-to-G editing activities and editing windows for Spy-and Nme2-ABE8e constructs in HEK293T cells. Numbers provided for each position in the protospacer represent the mean A-to-G editing efficiency across eight PAM-matched endogenous target sites, as measured via amplicon deep sequencing (n = 3 biological replicates). Crossed-out boxes indicate that no adenine was present at the specified position in the target panel tested.
Article Snippet: U6-driven sgRNA plasmids for the various Cas effectors were cloned using
Techniques: Construct, Plasmid Preparation, Transfection, Flow Cytometry, Amplification, Sequencing